Discussion
Enzymatic device
In this page, we firstly discuss the difference between the results of simulation and experiment. As shown in previous pages, the reaction velocity in simulation was faster than that in experiment. Disagreement of reaction kinetics between simulation and experiment seems to cause the difference. In particular, we referred to two articles [1,2] for the most of the rate constants for the simulation. We selected arbitrary but realistic rate constants for the rest. For an accurate simulation, fitting each rate constant to an experimental value is required.
In the simulation, single stranded DNAs are synthesized without limit because we assume constant concentration of dNTP. Since the concentration of dNTP is not constant in real experiment, the synthesis of single stranded DNAs will be saturated. Our model of the simulation can be refined if we take into account the saturation of Signal DNAs by formalizing Michaelis-Menten kinetics [3]. Moreover, our model ignores minor reactions such as denaturation of DNA. We can improve our model if we include such denaturing reaction.
In fact, we tried to fit the simulation results to experimental data by hand. We changed the rate constant of hybridization between Gate and Signal from 3.0×107[M-1min-1] to 1.0×105[M-1min-1], and that of other hybridization from 3.0×108[M-1min-1] to 3.0×106[M-1min-1]. We also added decrease terms to ordinary differential equations so that they converge to a constant value. The graph shown below is obtained by fitting.
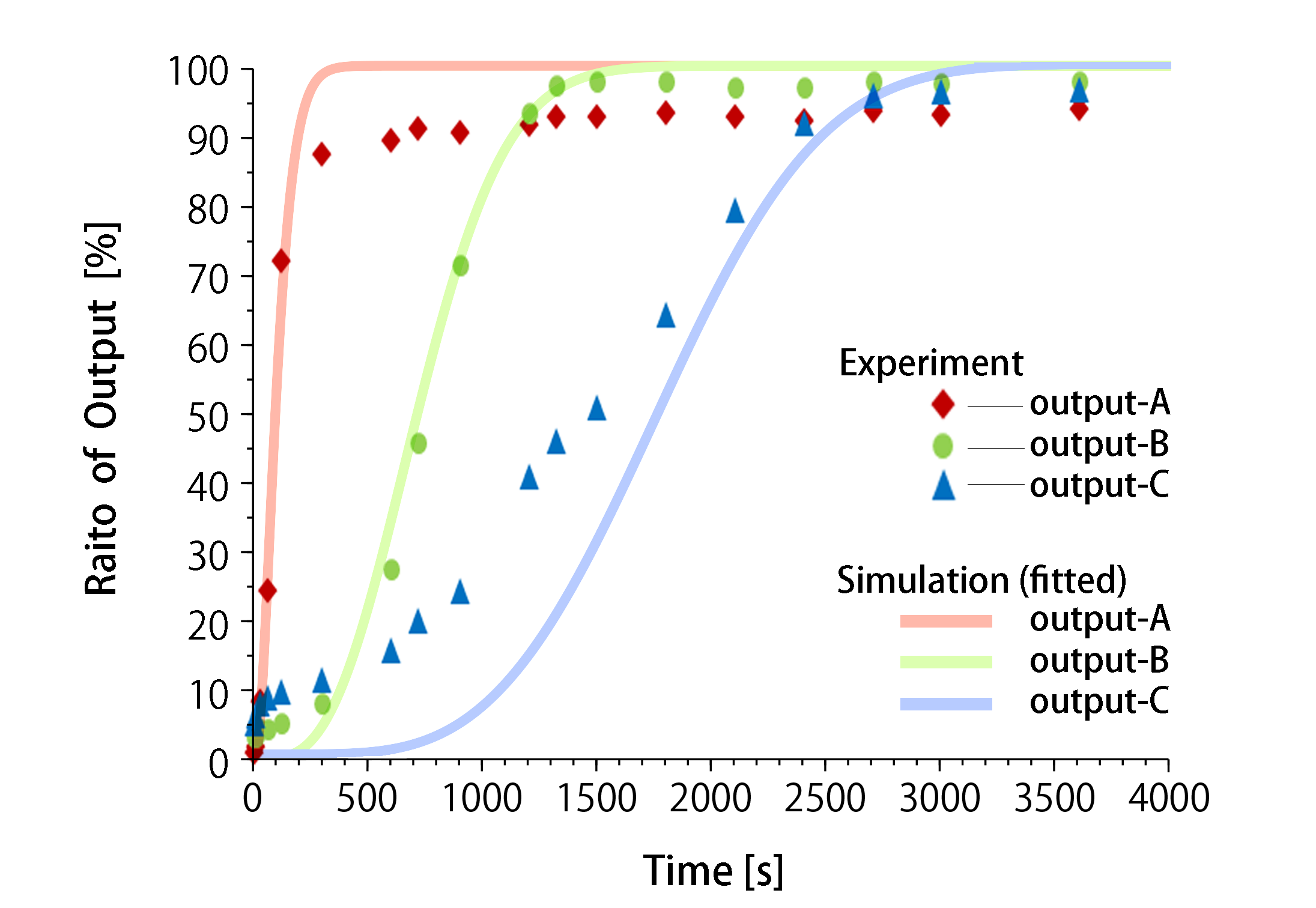
Fig.1 Fitting result of simulation (Enzyme dvice)
As described in the result of experiment, “ABC” or “CAB” can be released as output in order but the same output such as “ABB” or “ABA” cannot be released in the Enzymatic device. This is remained for future improvement.
Enzyme-free device
The outline of the graph of simulation and experiment differs from each other. We guess that insufficient thresholding caused the difference. The result of supplemental simulation without thresholding supports the idea (Fig.2). Concretely, the thresholding is caused by the difference of rate constant of two hybridizations: Signal to Output and Signal to Gate. The rate constant of them were the same in this simulation. Additionally, we tried to fit the simulation results to experimental data in the same way as Enzymatic decive. We changed the rate constant of seesawing reactions from 5.0×104[M-1s-1] to 1.0×104[M-1s-1].
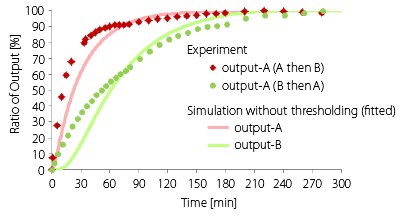
Fig.2 Fitting result of simulation (Enzyme-free dvice)
On the other hand, we may need to optimize the length of toehold to make experiment similar to the original simulation with thresholding. Purification of DNA structures is another possible solution because it can prevent stoichiometric inaccuracy. Such solutions may improve the performance of our Enzyme-free device in the future.
